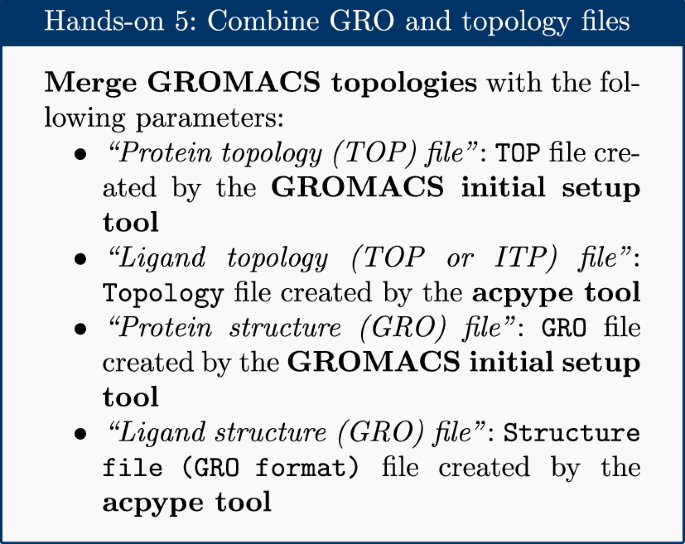
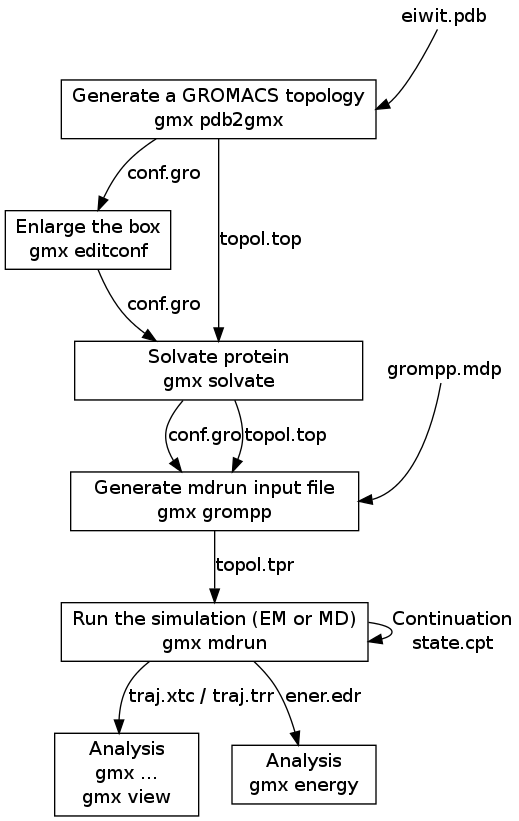
Intuitive, reproducible high-throughput molecular dynamics in Galaxy: a tutorial | Journal of Cheminformatics | Full Text

RTP File Extension | Gromacs Residue Topology Parameter File | Associated Programs | Free Online Tools - FileProInfo

Tutorial: Modelling post-translational modified proteins with GROMACS – Dr Anthony Nash MRSC: computational chemistry and life sciences

Data including GROMACS input files for atomistic molecular dynamics simulations of mixed, asymmetric bilayers including molecular topologies, equilibrated structures, and force field for lipids compatible with OPLS-AA parameters - ScienceDirect








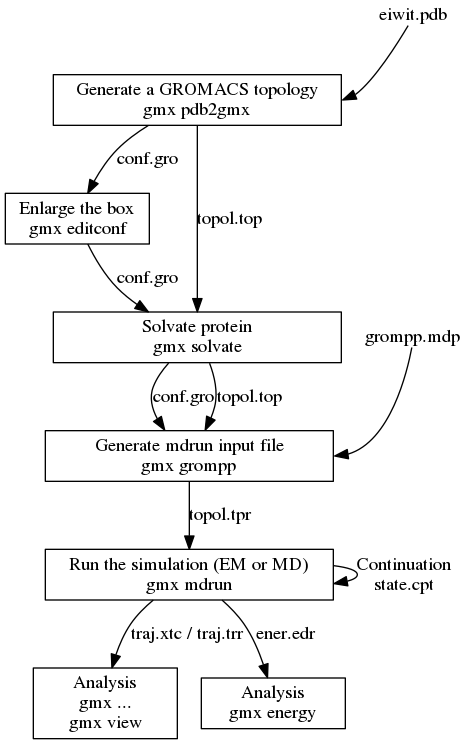

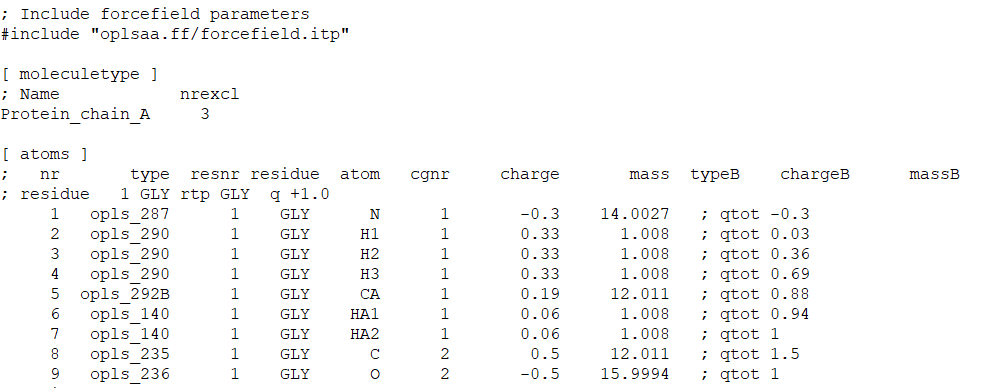

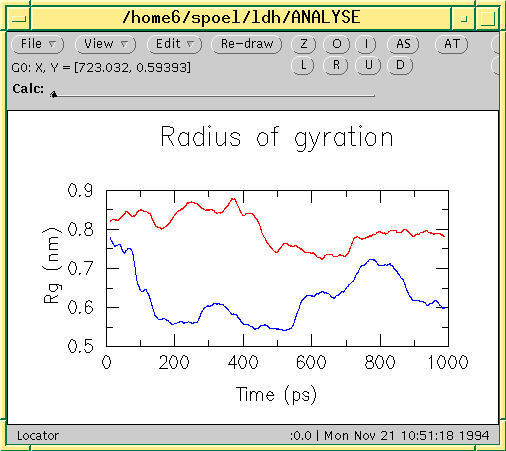


![PDF] GROMACS: A message-passing parallel molecular dynamics implementation | Semantic Scholar PDF] GROMACS: A message-passing parallel molecular dynamics implementation | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/625ddfe164e3a25405672ecc1ca17d05353ff182/8-Table1-1.png)